Stochastics
VIMC modelling groups produce stochastic data for us, which is currently 200 runs of their models varying key parameters, to express uncertainty. These files get uploaded to Box, and the groups are provided with guidance to produce files that we can process easily.
Stoner can process these stochastic files into a standard format to
make the files more predictable to work with. For a good compromise
between ease-of-use, performance and compression, we have used the
Parquet format and the arrow package. This
vignette covers how to process the data, how to create a central from a
set of stochastics, and how to see a graph of the stochastics for quick
diagnosis.
Creating standard stochastic files
This has superceded stoner_stochastic_process, which
will be removed and undocumented soon.
Here, we are mostly likely working with two folders or network
shares, one of which contains the stochastics that groups have uploaded,
and the other is a destination where we want to write our standardised
files. These I’ll call in_path and out_path.
The out_path folders I am assuming will be only written to
by stoner; paths and filenames will adhere to the following format:-
Within the root of the share we will have the touchstone, without a version. At time of writing, we have
202310gaviand202409malaria. I am not anticipating having separate stochastics for separate versions of the stochastics at this point.Within those touchstone folders, we have more folders in the form
Disease_Modelling-Group- such asCOVID_IC-GhaniorYF_IC-Garske. The groups and diseases set up here copy the historic contents of the Montagu database.Within each of these folders, we will be writing a file per country per scenario, containing all run_ids, named in the format
Modelling-Group_scenario_country.pq- for example,IC-Garske_yf-routine-ia2030_KEN.pq. If we also create a average (central) of the stochastics (see later), that will be calledIC-Garske_yf-routine-ia2030_central.pq, containing all the countries, and no run_id.
Example Usage - One file per scenario
Examples used so far are in the scripts folder in the
root of the stoner repo - files such as
process_stoch_202310gavi.R. Here is an example of the
Yellow Fever standardisation:-
scenarios = c("yf-no-vaccination", "yf-routine-bluesky",
"yf-routine-campaign-bluesky", "yf-routine-campaign-default",
"yf-routine-campaign-ia2030", "yf-routine-default",
"yf-routine-ia2030")
stoner::stone_stochastic_standardise(
group = "IC-Garske",
in_path = file.path(base_in_path, "YF-IC-Garske"),
out_path = file.path(base_out_path, "YF_IC-Garske"),
scenarios = scenarios,
files = "burden_results_stochastic_202310gavi-3_:scenario Keith Fraser.csv.xz")base_in_path and base_out_path are defined
elsewhere, and are my drive mappings to the incoming, and destination
network shares. We provide the group name, the paths to the uploaded
files (note the original dropbox path was a little incorrect with its
dashes instead of underscores!) - and the destination path in the format
described above.
We then provide the vector of scenarios, and because this group
follow the guidance very well, they have included the exact scenario in
the filenames. We can therefore use the :scenario glob in
the files argument, and stoner will handle each of the
seven scenarios in the right order.
Example Usage - One file per run
Other groups provide a file per run, which is also fine. Here, we can
specify an additional index parameter, usually
1:200, and use the glob :index in the files
argument. Here is an example of that - but note for this group, they
didn’t quite include the scenarios exactly in the filenames, so
we have to explicitly name the files, rather than using the
:scenario glob. This is still yellow fever, so using the
same scenarios vector as before.
stoner::stone_stochastic_standardise(
group = "UND-Perkins",
in_path = file.path(base_in_path, "YF-UND-Perkins"),
out_path = file.path(base_out_path, "YF_UND-Perkins"),
scenarios = scenarios,
files = c("stochastic_burden_est_YF_UND-Perkins_yf-no_vaccination_:index.csv.xz",
"stochastic_burden_est_YF_UND-Perkins_yf-routine_bluesky_:index.csv.xz",
"stochastic_burden_est_YF_UND-Perkins_yf-routine_campaign_bluesky_:index.csv.xz",
"stochastic_burden_est_YF_UND-Perkins_yf-routine_campaign_default_:index.csv.xz",
"stochastic_burden_est_YF_UND-Perkins_yf-routine_campaign_ia2030_:index.csv.xz",
"stochastic_burden_est_YF_UND-Perkins_yf-routine_default_:index.csv.xz",
"stochastic_burden_est_YF_UND-Perkins_yf-routine_ia2030_:index.csv.xz"),
index = 1:200)The subtle difference is that the group used underscores instead of
dashes in the scenario names. So here, I have to be careful and ensure
the order of files matches the order of
scenarios manually. After that, the :index
glob, and the index argument deal with the fact we have 200
files per scenario.
Example Usage - One file per country
If groups want to supply a file per country, then at present, we need
them to do so including the country in their uploads as normal, but
number their files using the index method above, rather
than having country names or ISO codes in the filename. We could do
better if this proves to be a popular method.
Advanced Usage
You shouldn’t have to worry about these things, but this is just so that you know they are there. We have some historic fixes that have made it possible for us to process some groups’ stochastics with some formatting issues, rather than asking them to resubmit. These fixes are enabled by default; they won’t do anything bad for groups where the fixes are not relevant, and are strictly required for the groups where they are relevant.
The Rubella Fix
An argument rubella_fix, by default TRUE,
deals with the fact that rubella groups have traditionally uploaded with
different outcome names. Instead of deaths,
rubella_deaths_congenital, and instead of
cases, rubella_cases_congenital. Some groups
additionally supplied the rubella_infections outcome.
The rubella_fix argument causes stoner to omit
rubella_infections, and rename the others to
deaths and cases respectively, standardising
them against the other diseases.
The Missing Run Id fix
One group also is in the habit of sending us one file per run_id, but
omitting the run_id field from their uploads. The argument
missing_run_id_fix, default TRUE lets Stoner
spot this, and specifically if index is 1:200
(ie, we are expecting 200 uploads), it will transplant the index into
the file to allow processing to be done normally.
Other Considerations:
Stoner rounds of all the outcomes, (deaths, cases, dalys, yll and cohort_size) to be integer-like, which can create some confusion with small countries with a very small number of cases, but we expect those confusions. Some groups still provide us with 16 decimal places, which creates some problems, and the integer-like rounding is in the guidance.
Note that for some groups with very large files (especially those with 16 decimal places), the processing takes a lot of memory - towards 128Gb. So run these on a large machine or perhaps a cluster node.
Creating a central estimate
For many groups, their central estimates are the average of their stochastic runs. Historically groups have provided their centrals before their stochastics, as a means of early review and detection of problems. Having standardised a group’s stochastic files, we can then generate an averaged central for a single touchstone, disease, group and scenario, with the following call:-
stone_stochastic_central(base, "202310gavi", "YF", "IC-Garske",
"yf-routine-campaign-ia2030")base here is the root of the share where our
standardised stochastic outputs have been put, and since those were
created by stoner, it knows how to build the paths correctly.
The result will be a new file in the form
group_scenario_central.pq so in this case
IC-Garske_yf-routine-campaign-ia2030_central.pq which
matches the other standardised filenames, except for the country; this
one central file will contain all countries.
The default averaging function is mean - if we really
want the median of the stochastics, then add the argument
avg_method = mean to the function call.
Creating stochastic graphs
It can be useful to quickly dig out a graph showing all the
stochastic lines together, to see what sort of spread they have. Again,
with base set to the root of the share where we are putting
our standardised outputs, here is an example function call, and the
graph that it produces.
stone_stochastic_graph(base, "202310gavi", "YF", "IC-Garske",
"KEN", "yf-routine-campaign-ia2030",
"deaths")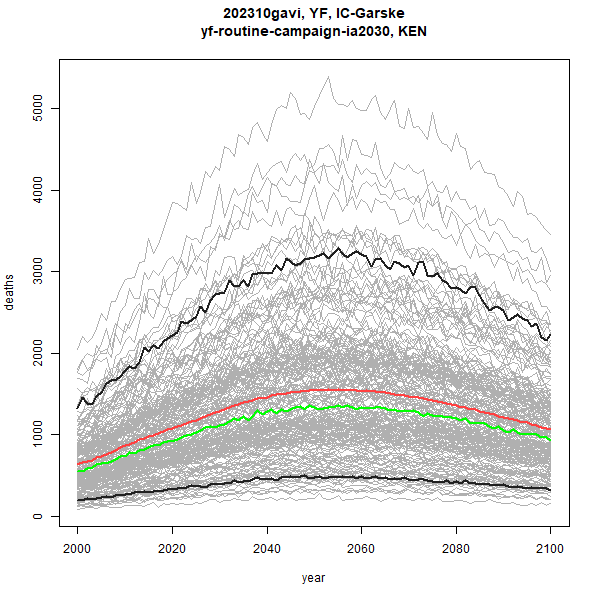
Red is the mean, and green is the median of the stochastics, and the
thicker black lines are the 5% and 95% quantiles. Other useful arguments
for graphs are by_cohort which plots a graph with birth
cohort on the x-axis instead of calendar year, log which
causes the y-axis to be log-scale, and ages can be a vector
of ages to filter to, for example 0:4 to plot stochastics
of under 5s.